-
MCX/MMC Suite - GPU-accelerated photon transport simulation platform
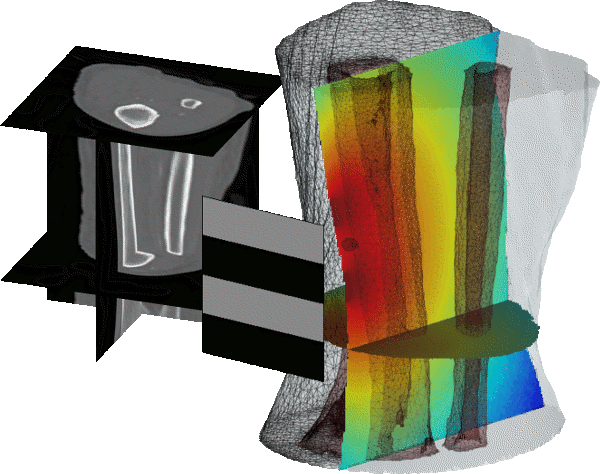
Monte Carlo eXtreme, or MCX, is an open-source Monte Carlo (MC) photon simulator for modeling light transport in 3D turbid media. The MCX/MMC Suite also contains mesh-based Monte Carlo (MMC) - an anatomically accurate 3D Monte Carlo (MC) simulator that can use a tetrahedral mesh to represent extremely complex tissue structures.
Both MCX and MMC support graphics processing units (GPUs) and are capable of simulating thousands of photons simultaneously, making them among the fastest and most physically-accurate photon modeling tools. Since their initial releases, the MCX Suite has become one of the most widely disseminated biophotonics modeling platforms, well-known for its high accuracy, high speed and versatility, as attested to by its over 27,000 downloads and over 1,100 citations from a large (2,400 registered users) world-wide user community. This rapidly expanding user community presents a diverse set of demands and a wide range of applications in the form of continuously evolving complex imaging domains and modalities. We are dedicated to serving this active and growing research community, and draw strong motivation from our extensive interactions with users, aiming to further expand the breadth, complexity, and efficiency of our simulation platform.
The development of MCX/MMC simulation platform is funded by the NIH/NIGMS under grant R01-GM114365.
Website http://mcx.space Wiki http://mcx.space/wiki Source code http://github.com/mcextreme User forum https://groups.google.com/g/mcx-users Download http://mcx.space/wiki/?Get Citation http://mcx.space/wiki/?Citation -
Iso2Mesh - 3D image based mesh generation toolbox
"Iso2Mesh" is a MATLAB/Octave-based mesh generation toolbox, designed for easy creation of high quality surface and tetrahedral meshes from 3D volumetric images. It contains over 200 mesh processing scripts/programs, working either independently or interacting with external free meshing utilities. Iso2Mesh toolbox can directly convert a 3D image stack, including binary, segmented or gray-scale images such as MRI or CT scans, into quality volumetric meshes. This makes it particularly suitable for multi-modality medical imaging data analysis and multi-physics modeling. Above all, iso2mesh is open-source. You can download it for free. You are also allowed to extend the toolbox for your own research and share with other users. Iso2Mesh is cross-platform and is compatible with both MATLAB and GNU Octave (a free MATLAB clone).
Between 2009 and 2020, Iso2Mesh has been cited over 620 times, with over 4,500 registered users worldwide, and an accumulative download over 40,000.
The details of this toolbox can be found in the following papers (citing the first paper is highly encouraged):
- - Anh Phong Tran, Shijie Yan and Qianqian Fang*, (2020) "Improving model-based fNIRS analysis using mesh-based anatomical and light-transport models," Neurophotonics, 7(1), 015008
- - Qianqian Fang and David Boas, "Tetrahedral mesh generation from volumetric binary and gray-scale images," Proceedings of IEEE International Symposium on Biomedical Imaging (ISBI 2009), pp. 1142-1145, 2009
Website http://iso2mesh.sf.net Wiki http://iso2mesh.sf.net Source code http://github.com/fangq/iso2mesh User forum https://groups.google.com/g/iso2mesh-users Download http://iso2mesh.sourceforge.net/cgi-bin/index.cgi?Download »Ubuntu/Debian sudo apt-get install octave-iso2meshDocumentation http://iso2mesh.sourceforge.net/cgi-bin/index.cgi?Doc »Fedora sudo dnf install octave-iso2meshIso2Mesh Suite
Currently, Iso2Mesh and its submodules are broadly distributed among popular open-source MATLAB toolboxes, especially among major neuroimaging tools, including
- - Fieldtrip (http://www.fieldtriptoolbox.org) [iso2mesh/jsonlab]
- - BrainStorm (https://neuroimage.usc.edu/brainstorm) [iso2mesh/brain2mesh/easyh5]
- - Lead-DBS (http://www.lead-dbs.org) [iso2mesh]
- - ROAST (https://www.parralab.org/roast) [iso2mesh]
- - HOMER2 (https://github.com/BUNPC/AtlasViewer) [iso2mesh/metch]
- - REST (https://github.com/goodshawn12/REST) [iso2mesh]
-
JSONLab - MATLAB JSON/UBJSON/MassagePack encoder/decoder
JSONLab is a free and open-source JSON/UBJSON/MessagePack encoder and decoder written in the native MATLAB language. It can be used to convert a MATLAB data structure (array, struct, cell, struct array, cell array, and objects) into JSON/UBJSON/MessagePack formatted strings and files, or to parse a JSON/UBJSON/MessagePack file into MATLAB data structure. JSONLab supports both MATLAB and GNU Octave (a free MATLAB clone).
JSON (JavaScript Object Notation) is a highly portable, human-readable and "fat-free" text format to represent complex and hierarchical data, widely used for data-exchange in applications. UBJSON (Universal Binary JSON) is a binary JSON format, specifically designed to specifically address the limitations of JSON, permitting efficient storage of binary data with strongly typed data records, resulting in smaller file sizes and fast encoding and decoding. MessagePack is another binary JSON-like data format widely used in data exchange in web/native applications. It is slightly more compact than UBJSON, but is not directly readable compared to UBJSON.
We envision that both JSON and its binary counterparts will play important rules not only for light-weight data storage, but also for storage and interchange of scientific data. It has both the flexibility and generality as in other general-purpose file specifications, such as HDF5 but has significantly reduced complexity and excellent readability.
Towards this goal, we have developed the JData Specification (http://github.com/fangq/jdata) to standardize serializations of complex scientific data structures, such as N-D arrays, sparse/complex-valued arrays, trees, maps, tables and graphs using JSON/binary JSON constructs. The text and binary formatted JData files are syntactically compatible with JSON/UBJSON formats, and can be readily parsed using existing JSON and UBJSON parsers. JSONLab is not just a parser and writer of JSON/UBJSON data files, but one that systematically converts complex scientific data structures into human-readable and universally supported JSON forms using the standardized JData data annotations.
JSONLab was named the Pick of the Week and Popular File 2018, ranking #6 in downloads among 35,500+ community-contributed toolboxes . Since its initial release, it has attracted over 47,000 downloads from MATLAB’s File Exchange and 14,000 from SourceForge.
Website http://openjdata.org/jsonlab MATLAB Central https://www.mathworks.com/matlabcentral/fileexchange/33381-jsonlab-a-toolbox-to-encode-decode-json-files Source code https://github.com/fangq/jsonlab »Debian sudo apt-get install octave-jsonlabUser forum https://groups.google.com/g/iso2mesh-users »Ubuntu sudo apt-get install octave-jsonlabDownload http://iso2mesh.sourceforge.net/cgi-bin/index.cgi?jsonlab/Download »Fedora sudo dnf install octave-jsonlabDocumentation http://iso2mesh.sourceforge.net/cgi-bin/index.cgi?jsonlab/Doc »Arch Linux sudo pikaur -S jsonlab -
OpenJData - Future-proof scientific data format
OpenJData is a set of open data-standards based on JSON/binary JSON formats. They are self-documenting, highly extensible, versatile, lightweight and easy-to-implement.
JData is a general-purpose data interchange format aimed for portability, readability and simplicity. It utilizes the JavaScript Object Notation (JSON) [RFC 4627] and Universal Binary JSON (UBJSON) specifications to store complex hierarchical data in both text and binary formats. In this specification, we define a list of JSON-compatible constructs to store a wide range of data structures, including scalars, arrays, structures, tables, hashes, linked lists, trees and graphs, and support optional data grouping and metadata for each data element. The generated data files are compatible with JSON/UBJSON specifications and can be readily processed by most existing parsers. Advanced features such as array compression, data linking and anchoring are supported to greatly enhance portability and scalability of the generated data files.
Website http://openjdata.org JData spec https://github.com/OpenJData/jdata Wiki http://openjdata.org/wiki BJData spec https://github.com/OpenJData/bjdata Source code https://github.com/OpenJData JNIfTI spec https://github.com/OpenJData/jnifti User forum https://github.com/fangq/jdata/issues JSNIRF spec https://github.com/OpenJData/jsnirf Download http://openjdata.org/wiki/#code JMesh spec https://github.com/OpenJData/jmesh Documentation http://openjdata.org/wiki Brain Mesh Library(in JNIfTI) https://github.com/OpenJData/BrainMeshLibrary MATLAB support http://openjdata.org/jsonlab
(source) http://github.com/fangq/jsonlab»Debian
»Fedorasudo apt-get install octave-jsonlabsudo dnf install octave-jsonlabPython support https://pypi.org/project/jdata/
(source) http://github.com/fangq/pyjdata»Debian
»pypisudo apt-get install pyjdatapip install jdata --userPython BJData support https://pypi.org/project/bjdata/
(source) http://github.com/fangq/pybj»Debian
»pypisudo apt-get install pybjpip install bjdata --userJavascript support https://github.com/fangq/jsdata -
Brain2mesh - One-liner brain mesh generator
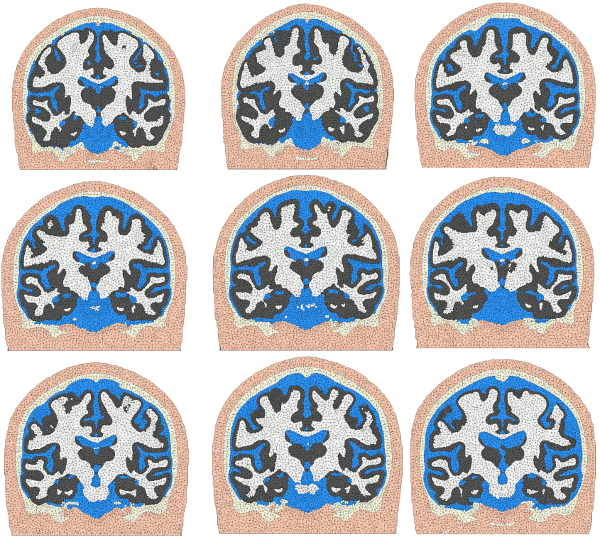
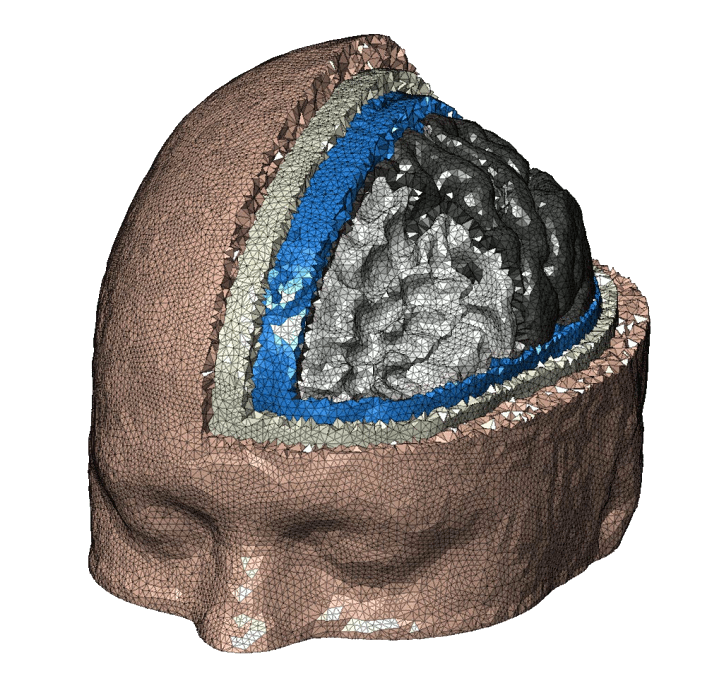
The Brain2Mesh toolbox provides a streamlined matlab function to convert a segmented brain volumes and surfaces into a high-quality multi-layered tetrahedral brain/full head mesh. Typical inputs include segmentation outputs from SPM, FreeSurfer, FSL etc. This tool does not handle the segmentation of MRI scans, but examples of how commonly encountered segmented datasets can be used to create meshes can be found in the package named brain2mesh-demos.
The details of this toolbox can be found in the following paper:
- Anh Phong Tran†, Shijie Yan†, and Qianqian Fang*, (2020) "Improving model-based fNIRS analysis using mesh-based anatomical and light-transport models," Neurophotonics, 7(1), 015008
Website http://mcx.space/brain2mesh Source code https://github.com/fangq/brain2mesh User forum https://groups.google.com/g/iso2mesh-users Download https://github.com/fangq/brain2mesh/releases/latest »Debian sudo apt-get install octave-brain2meshBrain mesh library https://github.com/OpenJData/BrainMeshLibrary »Fedora sudo dnf install octave-brain2mesh -
EasyH5 - Easy-to-use HDF5 data interface for MATLAB
EasyH5 is a fully automated, fast, compact and portable MATLAB object to HDF5 exporter/importer. It contains two easy-to-use functions -
loadh5.mandsaveh5.m. Thesaveh5.mcan handle almost all MATLAB data types, including structs, struct arrays, cells, cell arrays, real and complex arrays, strings, andcontainers.Mapobjects. All other data classes (such as a table, digraph, etc) can also be stored/loaded seemlessly using an undocumented data serialization interface (MATLAB only).EasyH5 stores complex numerical arrays using a special compound data type in an HDF5 dataset. The real-part of the data are stored as
Realand the imaginary part is stored as theImagcomponent. Theloadh5.mautomatically converts such data structure to a complex array. Starting from v0.8, EasyH5 also supports saving and loading sparse arrays using a compound dataset with 2 or 3 specialized subfields:SparseArray,Real, and, in the case of a sparse complex array,Imag. The sparse array dimension is stored as an attribute namedSparseArraySize, attached with the dataset. Using thedeflatefilter to save compressed arrays is supported in v0.8 and later.Website https://github.com/fangq/easyh5 MATLAB Central https://www.mathworks.com/matlabcentral/fileexchange/72918-easyh5 Source code https://github.com/fangq/easyh5 Download https://github.com/fangq/easyh5/releases/latest -
ZMat - A MATLAB/Octave data compression toolbox
ZMat provides both an easy-to-use C-based data compression library - libzmat as well a portable mex function to enable zlib/gzip/lzma/lzip/lz4/lz4hc based data compression/decompression and base64 encoding/decoding support in MATLAB and GNU Octave. It is fast and compact, can process a large array within a fraction of a second.
Among the 6 supported compression methods, lz4 is the fastest for compression/decompression; lzma is the slowest but has the highest compression ratio; zlib/gzip have the best balance between speed and compression time.
The ZMat MATLAB function accepts 3 types of inputs: char-based strings, numerical arrays or vectors, or logical arrays/vectors. Any other input format will result in an error unless you typecast the input into
int8/uint8format. A multi-dimensional numerical array is accepted, and the original input's type/dimension info is stored in the 2nd output"info". If one callszmatwith both the encoded data (in byte vector) and the"info"structure, zmat will first decode the binary data and then restore the original input's type and size.Website https://github.com/fangq/zmat MATLAB Central https://www.mathworks.com/matlabcentral/fileexchange/71434-zmat Source code https://github.com/fangq/zmat »Debian sudo apt-get install octave-zmat libzmat1-devDownload https://github.com/fangq/zmat/releases/latest »Fedora sudo dnf install octave-zmat zmat-devel -
JNIfTI Toolbox - Fast and portable NIfTI-1/2 reader and NIfTI-to-JNIfTI converter
This is a fully functional NIfTI-1/2 reader/writer that supports both MATLAB and GNU Octave, and is capable of reading/writing both non-compressed and compressed NIfTI files (
.nii, .nii.gz) as well as two-part Analyze7.5/NIfTI files (.hdr/.imgand.hdr.gz/.img.gz).More importantly, this is a toolbox that converts NIfTI data to its JSON-based replacement, JNIfTI (
.jniifor text-based and.bniifor binary-based), defined by the JNIfTI specification (http://github.com/fangq/jnifti). JNIfTI is a much more flexible, human-readable and extensible file format compared to the more rigid and opaque NIfTI format, making the data much easier to manipulate and share.The JNIfTI toolbox includes a stand-alone NIfTI-1/2 parser that works on both MATLAB and GNU Octave without needing additional components. To just reading and writing the un-compressed NIfTI and Analyze7.5 files (.nii, .hdr/.img), one only needs to run
addpath('/path/to/jnifti'). For MATLAB, JNIfTI toolbox utilizesmemmapfile-based disk-reading, making it very fast. For Octave,memmapfileis currently not implemented, so, a full reading is required.The JNIfTI toolbox is also capable of reading/writing gzip-compressed NIfTI and Analyze7.5 files (
.nii.gz, .hdr.gz, .img.gz). This feature is supported in MATLAB directly without needing another toolbox (MATLAB must be in the JVM-enabled mode).To process gzip-compressed NIfTI/Analyze files in Octave and MATLAB with
-nojvm, one need to install the open-source JSONLab and ZMat toolboxes, both supporting MATLAB and Octave.To save NIfTI-1/2 data as JNIfTI files, one needs to install JSONLab. The JNIfTI data format supports internal compression (as oppose to external compression such as
*.gzfiles). To create or read compressed JNIfTI files in Octave, one must install the ZMat toolbox, as listed above.Website https://github.com/fangq/jnifti/tree/master/lib/matlab MATLAB Central https://www.mathworks.com/matlabcentral/fileexchange/72756-jnifti Source code https://github.com/fangq/jnifti/tree/master/lib/matlab »Debian sudo apt-get install octave-jniftiDownload https://github.com/fangq/jnifti/releases/latest »Fedora sudo dnf install octave-jnifti
-
 MCX/MMC
MCX/MMC
-
 Iso2Mesh
Iso2Mesh
-
 JSONLab
JSONLab
-
 OpenJData
OpenJData
-
 Brain2Mesh
Brain2Mesh
-
 EasyH5
EasyH5
-
 ZMat
ZMat
-
 JNIfTI
JNIfTI
![[Home]](upload/lab_full_logo.png)